|
| class | AAIndex |
| | Representation of selected AAIndex properties. More...
|
| |
| class | AASequence |
| | Representation of a peptide/protein sequence. More...
|
| |
| class | AbsoluteQuantitation |
| | AbsoluteQuantitation is a class to support absolute or relative quantitation for targeted or untargeted quantitation workflows (e.g., Isotope Dilution Mass Spectrometry). More...
|
| |
| class | AbsoluteQuantitationMethod |
| | AbsoluteQuantitationMethod is a class to hold information about the quantitation method and for applying and/or generating the quantitation method. More...
|
| |
| class | AbsoluteQuantitationMethodFile |
| | File adapter for AbsoluteQuantitationMethod files. More...
|
| |
| class | AbsoluteQuantitationStandards |
| | AbsoluteQuantitationStandards is a class to handle the relationship between runs, components, and their actual concentrations. More...
|
| |
| class | AbsoluteQuantitationStandardsFile |
| | Load files containing runConcentration data. More...
|
| |
| class | AccurateMassSearchEngine |
| | An algorithm to search for exact mass matches from a spectrum against a database (e.g. HMDB). More...
|
| |
| class | AccurateMassSearchResult |
| |
| class | Acquisition |
| | Information about one raw data spectrum that was combined with several other raw data spectra. More...
|
| |
| class | AcquisitionInfo |
| | Description of the combination of raw data to a single spectrum. More...
|
| |
| class | AcquisitionInfoVisualizer |
| | Class that displays all meta information for AcquisitionInfo objects. More...
|
| |
| class | AcquisitionVisualizer |
| | Class that displays all meta information for Acquisition objects. More...
|
| |
| class | Adduct |
| |
| class | AdductInfo |
| |
| class | AhoCorasickAmbiguous |
| | Extended Aho-Corasick algorithm capable of matching ambiguous amino acids in the pattern (i.e. proteins). More...
|
| |
| class | Annotation1DCaret |
| | An annotation item which paints a set of carets on the canvas. More...
|
| |
| class | Annotation1DDistanceItem |
| | An annotation item which represents a measured distance between two peaks. More...
|
| |
| class | Annotation1DItem |
| | An abstract class acting as an interface for the different 1D annotation items. More...
|
| |
| class | Annotation1DPeakItem |
| | A peak annotation item. More...
|
| |
| class | Annotation1DTextItem |
| | An annotation item which represents an arbitrary text on the canvas. More...
|
| |
| class | Annotations1DContainer |
| | Container for annotations to content of Spectrum1DCanvas. More...
|
| |
| struct | AnnotationStatistics |
| |
| class | AScore |
| | Implementation of the Ascore For a given peptide sequence and its MS/MS spectrum it identifies the most probable phosphorylation-site(s). For each phosphorylation site a probability score is calculated. The algorithm is implemented according to Beausoleil et al. (Nat. Biotechnol. 2006). More...
|
| |
| class | AverageLinkage |
| | AverageLinkage ClusterMethod. More...
|
| |
| class | AxisPainter |
| | Draws a coordinate axis. It has only static methods, that's why the constructor is private. More...
|
| |
| class | AxisTickCalculator |
| | Calculates ticks for a given value range. More...
|
| |
| class | AxisWidget |
| | Widget that represents an axis of a graph. More...
|
| |
| class | BackgroundControl |
| |
| class | BackgroundIntensityBin |
| |
| class | Base64 |
| | Class to encode and decode Base64. More...
|
| |
| class | BaseFeature |
| | A basic LC-MS feature. More...
|
| |
| class | BaseGroupFinder |
| | The base class of all element group finding algorithms. More...
|
| |
| class | BaseLabeler |
| | Abstract base class for all kinds of labeling techniques. More...
|
| |
| class | BaseModel |
| | Abstract base class for all D-dimensional models. More...
|
| |
| class | BaseSuperimposer |
| | The base class of all superimposer algorithms. More...
|
| |
| class | BaseVisualizer |
| | A base class for all visualizer classes. More...
|
| |
| class | BaseVisualizerGUI |
| | A base class for all visualizer classes. More...
|
| |
| class | BasicProteinInferenceAlgorithm |
| | Algorithm class that implements simple protein inference by aggregation of peptide scores. It has multiple parameter options like the aggregation method, when to distinguish peptidoforms, and if you want to use shared peptides ("use_shared_peptides"). First, the best PSM per spectrum is used, then only the best PSM per peptidoform is aggregated. Peptidoforms can optionally be distinguished via the treat_X_separate parameters: More...
|
| |
| class | BayesianProteinInferenceAlgorithm |
| | Performs a Bayesian protein inference on Protein/Peptide identifications or ConsensusMap (experimental). More...
|
| |
| class | BernNorm |
| | BernNorm scales the peaks by ranking them and then scaling them according to rank. More...
|
| |
| class | BiGaussFitter1D |
| | BiGaussian distribution fitter (1-dim.) approximated using linear interpolation. More...
|
| |
| class | BiGaussModel |
| | BiGaussian distribution approximated using linear interpolation. More...
|
| |
| class | BinaryTreeNode |
| | Elements of a binary tree used to represent a hierarchical clustering process. More...
|
| |
| class | BinnedSharedPeakCount |
| | Compare functor scoring the shared peaks for similarity measurement. More...
|
| |
| class | BinnedSpectralContrastAngle |
| | Compare functor scoring the spectral contrast angle for similarity measurement. More...
|
| |
| class | BinnedSpectrum |
| | This is a binned representation of a PeakSpectrum. More...
|
| |
| class | BinnedSpectrumCompareFunctor |
| | Base class for compare functors of BinnedSpectra. More...
|
| |
| class | BinnedSumAgreeingIntensities |
| | Sum of agreeing intensities for similarity measurement. More...
|
| |
| class | BSpline2d |
| | b spline interpolation More...
|
| |
| class | Bzip2Ifstream |
| | Decompresses files which are compressed in the bzip2 format (*.bz2) More...
|
| |
| class | Bzip2InputStream |
| | Implements the BinInputStream class of the xerces-c library in order to read bzip2 compressed XML files. More...
|
| |
| class | CachedmzML |
| | An class that uses on-disk caching to read and write spectra and chromatograms. More...
|
| |
| class | CachedSwathFileConsumer |
| | On-disk cached implementation of FullSwathFileConsumer. More...
|
| |
| class | CalibrationData |
| | A helper class, holding all calibration points. More...
|
| |
| class | CentroidData |
| |
| class | CentroidPeak |
| |
| class | ChargePair |
| | Representation of a (putative) link between two Features, which stem from the same compound but have different charge (including different adduct ions (H+, Na+, ..) More...
|
| |
| class | ChromatogramExtractor |
| | The ChromatogramExtractor extracts chromatograms from a spectra file. More...
|
| |
| class | ChromatogramExtractorAlgorithm |
| | The ChromatogramExtractorAlgorithm extracts chromatograms from a MS data. More...
|
| |
| class | ChromatogramPeak |
| | A 1-dimensional raw data point or peak for chromatograms. More...
|
| |
| class | ChromatogramSettings |
| | Representation of chromatogram settings, e.g. SRM/MRM chromatograms. More...
|
| |
| class | ChromatogramTools |
| | Conversion class to convert chromatograms. More...
|
| |
| class | ChromeleonFile |
| | Load Chromeleon HPLC text file and save it into a `MSExperiment`. More...
|
| |
| struct | Citation |
| | Stores Citations for individual TOPP tools. More...
|
| |
| class | ClusterAnalyzer |
| | Bundles analyzing tools for a clustering (given as sequence of BinaryTreeNode's) More...
|
| |
| class | ClusteredMS2ConsensusSpectrum |
| |
| class | ClusterFunctor |
| | Base class for cluster functors. More...
|
| |
| class | ClusterHierarchical |
| | Hierarchical clustering with generic clustering functions. More...
|
| |
| class | ClusteringGrid |
| | data structure to store 2D data to be clustered e.g. (m/z, retention time) coordinates from multiplex filtering More...
|
| |
| class | ClusterProxyKD |
| | Proxy for a (potential) cluster. More...
|
| |
| class | CmpHypothesesByScore |
| |
| class | CmpMassTraceByMZ |
| |
| class | CoarseIsotopePatternGenerator |
| | Isotope pattern generator for coarse isotope distributions. More...
|
| |
| class | ColorSelector |
| | A widget for selecting a color. More...
|
| |
| class | ComplementFilter |
| | total intensity of peak pairs that could result from complementing fragments of charge state 1 More...
|
| |
| class | ComplementMarker |
| | ComplementMarker marks peak pairs which could represent y - b ion pairs. More...
|
| |
| class | CompleteLinkage |
| | CompleteLinkage ClusterMethod. More...
|
| |
| class | CompNovoIdentification |
| | run with CompNovoIdentification More...
|
| |
| class | CompNovoIdentificationBase |
| | run with CompNovoIdentificationBase More...
|
| |
| class | CompNovoIdentificationCID |
| | run with CompNovoIdentificationCID More...
|
| |
| class | CompNovoIonScoring |
| | run with CompNovoIonScoring More...
|
| |
| class | CompNovoIonScoringBase |
| | run with CompNovoIonScoringBase More...
|
| |
| class | CompNovoIonScoringCID |
| | run with CompNovoIonScoringCID More...
|
| |
| class | Compomer |
| | Holds information on an edge connecting two features from a (putative) charge ladder. More...
|
| |
| class | CompressedInputSource |
| | This class is based on xercesc::LocalFileInputSource. More...
|
| |
| class | ConfidenceScoring |
| |
| struct | ConnectedComponent |
| |
| class | ConsensusFeature |
| | A consensus feature spanning multiple LC-MS/MS experiments. More...
|
| |
| class | ConsensusIDAlgorithm |
| | Abstract base class for all ConsensusID algorithms (that calculate a consensus from multiple ID runs). More...
|
| |
| class | ConsensusIDAlgorithmAverage |
| | Calculates a consensus from multiple ID runs by averaging the search scores. More...
|
| |
| class | ConsensusIDAlgorithmBest |
| | Calculates a consensus from multiple ID runs by taking the best search score. More...
|
| |
| class | ConsensusIDAlgorithmIdentity |
| | Abstract base class for ConsensusID algorithms that compare only identical sequences. More...
|
| |
| class | ConsensusIDAlgorithmPEPIons |
| | Calculates a consensus from multiple ID runs based on PEPs and shared ions. More...
|
| |
| class | ConsensusIDAlgorithmPEPMatrix |
| | Calculates a consensus from multiple ID runs based on PEPs and sequence similarities. More...
|
| |
| class | ConsensusIDAlgorithmRanks |
| | Calculates a consensus from multiple ID runs based on the ranks of the search hits. More...
|
| |
| class | ConsensusIDAlgorithmSimilarity |
| | Abstract base class for ConsensusID algorithms that take peptide similarity into account. More...
|
| |
| class | ConsensusIDAlgorithmWorst |
| | Calculates a consensus from multiple ID runs by taking the worst search score (conservative approach). More...
|
| |
| class | ConsensusIsotopePattern |
| |
| class | ConsensusMap |
| | A container for consensus elements. More...
|
| |
| class | ConsensusMapMergerAlgorithm |
| | Merges identification data in ConsensusMaps. More...
|
| |
| class | ConsensusMapNormalizerAlgorithmMedian |
| | Algorithms of ConsensusMapNormalizer. More...
|
| |
| class | ConsensusMapNormalizerAlgorithmQuantile |
| | Algorithms of ConsensusMapNormalizer. More...
|
| |
| class | ConsensusMapNormalizerAlgorithmThreshold |
| | Algorithms of ConsensusMapNormalizer. More...
|
| |
| class | ConsensusXMLFile |
| | This class provides Input functionality for ConsensusMaps and Output functionality for alignments and quantitation. More...
|
| |
| class | ConsoleUtils |
| |
| class | ConstRefVector |
| | This vector holds pointer to the elements of another container. More...
|
| |
| class | ContactPerson |
| | Contact person information. More...
|
| |
| class | ContactPersonVisualizer |
| | Class that displays all meta information for ContactPerson objects. More...
|
| |
| class | Contaminants |
| | This class is a metric for the QualityControl TOPP tool. More...
|
| |
| class | ContinuousWaveletTransform |
| | This class is the base class of the continuous wavelet transformation. More...
|
| |
| class | ContinuousWaveletTransformNumIntegration |
| | This class computes the continuous wavelet transformation using a marr wavelet. More...
|
| |
| class | ControlledVocabulary |
| | Representation of a controlled vocabulary. More...
|
| |
| class | ConvexHull2D |
| | A 2-dimensional hull representation in [counter]clockwise direction - depending on axis labelling. More...
|
| |
| class | CrossLinksDB |
| |
| class | CsiFingerIdMzTabWriter |
| |
| class | CsvFile |
| | This class handles csv files. Currently only loading is implemented. More...
|
| |
| class | CubicSpline2d |
| | cubic spline interpolation as described in R.L. Burden, J.D. Faires, Numerical Analysis, 4th ed. PWS-Kent, 1989, ISBN 0-53491-585-X, pp. 126-131. More...
|
| |
| class | CVMappingFile |
| | Used to load CvMapping files. More...
|
| |
| class | CVMappingRule |
| | Representation of a CV Mapping rule used by CVMappings. More...
|
| |
| class | CVMappings |
| | Representation of controlled vocabulary mapping rules (for PSI formats) More...
|
| |
| class | CVMappingTerm |
| | Representation of controlled vocabulary term. More...
|
| |
| class | CVReference |
| | Controlled Vocabulary Reference. More...
|
| |
| class | CVTerm |
| | Representation of controlled vocabulary term. More...
|
| |
| class | CVTermList |
| | Representation of controlled vocabulary term list. More...
|
| |
| class | CVTermListInterface |
| | Interface to the controlled vocabulary term list. More...
|
| |
| class | DataFilterDialog |
| | Dialog for creating and changing a DataFilter. More...
|
| |
| class | DataFilters |
| | DataFilter array providing some convenience functions. More...
|
| |
| class | DataProcessing |
| | Description of the applied preprocessing steps. More...
|
| |
| class | DataProcessingVisualizer |
| | Class that displays all meta information for DataProcessing objects. More...
|
| |
| class | DataValue |
| | Class to hold strings, numeric values, lists of strings and lists of numeric values. More...
|
| |
| class | Date |
| | Date Class. More...
|
| |
| class | DateTime |
| | DateTime Class. More...
|
| |
| struct | DaTrait |
| |
| class | DBoundingBox |
| | A D-dimensional bounding box. More...
|
| |
| class | DeconvPeak |
| |
| class | DecoyHelper |
| | Helper class for calculcations on decoy proteins. More...
|
| |
| class | DefaultParamHandler |
| | A base class for all classes handling default parameters. More...
|
| |
| class | Deisotoper |
| |
| class | DeNovoAlgorithm |
| | Base class for ion scoring implementation for de novo algorithms. More...
|
| |
| class | DeNovoIdentification |
| | Base class for de novo identification. More...
|
| |
| class | DeNovoIonScoring |
| | Base class for ion scoring implementation for de novo algorithms. More...
|
| |
| class | DeNovoPostScoring |
| | Base class for ion scoring implementation for de novo algorithms. More...
|
| |
| class | DetectabilitySimulation |
| | Simulates peptide detectability. More...
|
| |
| class | DiaPrescore |
| | Scoring of an spectrum given library intensities of a transition group. More...
|
| |
| class | DIAScoring |
| | Scoring of an spectrum at the peak apex of an chromatographic elution peak. More...
|
| |
| class | Digestion |
| | Meta information about digestion of a sample. More...
|
| |
| class | DigestionEnzyme |
| | Abstract base class for digestion enzymes. More...
|
| |
| class | DigestionEnzymeDB |
| | Digestion enzyme database (base class) More...
|
| |
| class | DigestionEnzymeProtein |
| | Representation of a digestion enzyme for proteins (protease) More...
|
| |
| class | DigestionEnzymeRNA |
| | Representation of a digestion enzyme for RNA (RNase) More...
|
| |
| class | DigestionVisualizer |
| | Class that displays all meta information of digestion objects. More...
|
| |
| class | DigestSimulation |
| | Simulates protein digestion. More...
|
| |
| class | DistanceMatrix |
| | A two-dimensional distance matrix, similar to OpenMS::Matrix. More...
|
| |
| class | DocumentIdentifier |
| | Manage source document information. More...
|
| |
| class | DocumentIdentifierVisualizer |
| | Class that displays all meta information for DocumentIdentifier objects. More...
|
| |
| class | DocumentIDTagger |
| | Tags OpenMS file containers with a DocumentID. More...
|
| |
| struct | DPeak |
| | Metafunction to choose among Peak1D respectively Peak2D through a template argument. More...
|
| |
| class | DPosition |
| | Representation of a coordinate in D-dimensional space. More...
|
| |
| class | DRange |
| | A D-dimensional half-open interval. More...
|
| |
| class | DTA2DFile |
| | DTA2D File adapter. More...
|
| |
| class | DTAFile |
| | File adapter for DTA files. More...
|
| |
| class | EDTAFile |
| | File adapter for Enhanced DTA files. More...
|
| |
| class | EGHFitter1D |
| | Exponential-Gaussian hybrid distribution fitter (1-dim.) using Levenberg-Marquardt algorithm (Eigen implementation) for parameter optimization. More...
|
| |
| class | EGHModel |
| | Exponential-Gaussian hybrid distribution model for elution profiles. More...
|
| |
| class | EGHTraceFitter |
| | A RT Profile fitter using an Exponential Gaussian Hybrid background model. More...
|
| |
| class | Element |
| | Representation of an element. More...
|
| |
| class | ElementDB |
| | Stores elements. More...
|
| |
| class | ElutionModelFitter |
| | Helper class for fitting elution models to features. More...
|
| |
| class | ElutionPeakDetection |
| | Extracts chromatographic peaks from a mass trace. More...
|
| |
| class | EmgFitter1D |
| | Exponentially modified gaussian distribution fitter (1-dim.) using Levenberg-Marquardt algorithm (Eigen implementation) for parameter optimization. More...
|
| |
| class | EmgGradientDescent |
| | Compute the area, background and shape metrics of a peak. More...
|
| |
| class | EmgGradientDescent_friend |
| |
| class | EmgModel |
| | Exponentially modified gaussian distribution model for elution profiles. More...
|
| |
| class | EmgScoring |
| | Scoring of an elution peak using an exponentially modified gaussian distribution model. More...
|
| |
| class | EmpiricalFormula |
| | Representation of an empirical formula. More...
|
| |
| class | EnhancedTabBar |
| | Convenience tab bar implementation. More...
|
| |
| class | EnhancedTabBarWidgetInterface |
| | Widgets that are placed into an EnhancedTabBar must implement this interface. More...
|
| |
| class | EnhancedWorkspace |
| |
| class | EnzymaticDigestion |
| | Class for the enzymatic digestion of sequences. More...
|
| |
| class | EnzymaticDigestionLogModel |
| | Class for the Log L model of enzymatic digestion of proteins. More...
|
| |
| struct | EqualInTolerance |
| | Struct for binary predicate to consider equality with a certain tolerance. More...
|
| |
| class | EuclideanSimilarity |
| | CompareFunctor for 2Dpoints. More...
|
| |
| class | ExperimentalDesign |
| | Representation of the Experimental Design in OpenMS. Instances can be loaded via the ExperimentalDesignFile class. More...
|
| |
| class | ExperimentalDesignFile |
| | Provides means to load an ExperimentalDesign from a TSV file. More...
|
| |
| class | ExperimentalSettings |
| | Description of the experimental settings. More...
|
| |
| class | ExperimentalSettingsVisualizer |
| | Class that displays all meta information for ExperimentalSettings objects. More...
|
| |
| class | ExtendedIsotopeFitter1D |
| | Extended isotope distribution fitter (1-dim.) approximated using linear interpolation. More...
|
| |
| class | ExtendedIsotopeModel |
| | Extended isotope distribution approximated using linear interpolation. More...
|
| |
| class | Factory |
| | Returns FactoryProduct* based on the name of the desired concrete FactoryProduct. More...
|
| |
| class | FactoryBase |
| | Base class for Factory<T> More...
|
| |
| class | FakeProcess |
| | A FakeProcess class. More...
|
| |
| class | FalseDiscoveryRate |
| | Calculates false discovery rates (FDR) from identifications. More...
|
| |
| class | FASTAContainer |
| | template parameter for vector-based FASTA access More...
|
| |
| class | FASTAContainer< TFI_File > |
| | FASTAContainer<TFI_File> will make FASTA entries available chunk-wise from start to end by loading it from a FASTA file. This avoids having to load the full file into memory. While loading, the container will memorize the file offsets of each entry, allowing to read an arbitrary i'th entry again from disk. If possible, only entries from the currently cached chunk should be queried, otherwise access will be slow. More...
|
| |
| class | FASTAContainer< TFI_Vector > |
| | FASTAContainer<TFI_Vector> simply takes an existing vector of FASTAEntries and provides the same interface with a potentially huge speed benefit over FASTAContainer<TFI_File> since it does not need disk access, but at the cost of memory. More...
|
| |
| class | FASTAFile |
| | This class serves for reading in and writing FASTA files. More...
|
| |
| class | Feature |
| | An LC-MS feature. More...
|
| |
| class | FeatureDeconvolution |
| | An algorithm to decharge features (i.e. as found by FeatureFinder). More...
|
| |
| class | FeatureDistance |
| | A functor class for the calculation of distances between features or consensus features. More...
|
| |
| class | FeatureEditDialog |
| | Dialog for editing a feature. More...
|
| |
| class | FeatureFileOptions |
| | Options for loading files containing features. More...
|
| |
| class | FeatureFinder |
| | The main feature finder class. More...
|
| |
| class | FeatureFinderAlgorithm |
| | Abstract base class for FeatureFinder algorithms. More...
|
| |
| class | FeatureFinderAlgorithmIsotopeWavelet |
| | Implements the isotope wavelet feature finder. More...
|
| |
| class | FeatureFinderAlgorithmMRM |
| | FeatureFinderAlgorithm for MRM experiments. More...
|
| |
| class | FeatureFinderAlgorithmPicked |
| | FeatureFinderAlgorithm for picked peaks. More...
|
| |
| struct | FeatureFinderAlgorithmPickedHelperStructs |
| | Wrapper struct for all the classes needed by the FeatureFinderAlgorithmPicked and the associated classes. More...
|
| |
| class | FeatureFinderAlgorithmSH |
| | The Superhirn FeatureFinderAlgorithm. More...
|
| |
| class | FeatureFinderAlgorithmSHCtrl |
| | A facade for various Superhirn FeatureFinder classes. Use FeatureFinderAlgorithmSH instead. More...
|
| |
| struct | FeatureFinderDefs |
| | The purpose of this struct is to provide definitions of classes and typedefs which are used throughout all FeatureFinder classes. More...
|
| |
| class | FeatureFinderIdentificationAlgorithm |
| |
| class | FeatureFinderMultiplexAlgorithm |
| |
| class | FeatureFindingMetabo |
| | Method for the assembly of mass traces belonging to the same isotope pattern, i.e., that are compatible in retention times, mass-to-charge ratios, and isotope abundances. More...
|
| |
| class | FeatureGroupingAlgorithm |
| | Base class for all feature grouping algorithms. More...
|
| |
| class | FeatureGroupingAlgorithmKD |
| | A feature grouping algorithm for unlabeled data. More...
|
| |
| class | FeatureGroupingAlgorithmLabeled |
| | A map feature grouping algorithm for labeling techniques with two labels. More...
|
| |
| class | FeatureGroupingAlgorithmQT |
| | A feature grouping algorithm for unlabeled data. More...
|
| |
| class | FeatureGroupingAlgorithmUnlabeled |
| | A map feature grouping algorithm for unlabeled data. More...
|
| |
| class | FeatureHandle |
| | Representation of a Peak2D, RichPeak2D or Feature . More...
|
| |
| class | FeatureHypothesis |
| | Internal structure used in FeatureFindingMetabo that keeps track of a feature hypothesis (isotope group hypothesis). More...
|
| |
| class | FeatureLCProfile |
| |
| class | FeatureMap |
| | A container for features. More...
|
| |
| class | FeatureMapping |
| |
| class | FeatureOpenMS |
| | An implementation of the OpenSWATH Feature Access interface using OpenMS. More...
|
| |
| class | FeatureXMLFile |
| | This class provides Input/Output functionality for feature maps. More...
|
| |
| class | FIAMSDataProcessor |
| | Data processing for FIA-MS data. More...
|
| |
| class | FIAMSScheduler |
| |
| class | File |
| | Basic file handling operations. More...
|
| |
| class | FileHandler |
| | Facilitates file handling by file type recognition. More...
|
| |
| struct | FileTypes |
| | Centralizes the file types recognized by FileHandler. More...
|
| |
| class | FileWatcher |
| | Watcher that monitors file changes. More...
|
| |
| class | FilterFunctor |
| | A FilterFunctor extracts some spectrum characteristics for quality assessment. More...
|
| |
| class | FineIsotopePatternGenerator |
| | Isotope pattern generator for fine isotope distributions. More...
|
| |
| class | Fitter1D |
| | Abstract base class for all 1D-dimensional model fitter. More...
|
| |
| class | FragmentMassError |
| |
| class | FTPeakDetectController |
| |
| class | FullSwathFileConsumer |
| | Abstract base class which can consume spectra coming from SWATH experiment stored in a single file. More...
|
| |
| class | FuzzyStringComparator |
| | Fuzzy comparison of strings, tolerates numeric differences. More...
|
| |
| class | GaussFilter |
| | This class represents a Gaussian lowpass-filter which works on uniform as well as on non-uniform profile data. More...
|
| |
| class | GaussFilterAlgorithm |
| | This class represents a Gaussian lowpass-filter which works on uniform as well as on non-uniform profile data. More...
|
| |
| class | GaussFitter1D |
| | Gaussian distribution fitter (1-dim.) approximated using linear interpolation. More...
|
| |
| class | GaussModel |
| | Normal distribution approximated using linear interpolation. More...
|
| |
| class | GaussTraceFitter |
| | Fitter for RT profiles using a Gaussian background model. More...
|
| |
| class | GoodDiffFilter |
| | GoodDiffFilter counts the number ob peak pairs whose m/z difference can be explained by a amino acid loss. More...
|
| |
| class | Gradient |
| | Representation of a HPLC gradient. More...
|
| |
| class | GradientVisualizer |
| | GradientVisualizer is a visualizer class for objects of type gradient. More...
|
| |
| class | GridBasedCluster |
| | basic data structure for clustering More...
|
| |
| class | GridBasedClustering |
| | 2D hierarchical clustering implementation optimized for large data sets containing many small clusters i.e. dimensions of clusters << dimension of entire dataset More...
|
| |
| class | GridFeature |
| | Representation of a feature in a hash grid. More...
|
| |
| class | GridSearch |
| |
| class | GUIHelpers |
| | Class which holds static GUI-related helper functions. More...
|
| |
| class | GUIProgressLoggerImpl |
| | Implements a GUI version of the ProgressLoggerImpl. More...
|
| |
| class | GzipIfstream |
| | Decompresses files which are compressed in the gzip format (*.gzip) More...
|
| |
| class | GzipInputStream |
| | Implements the BinInputStream class of the xerces-c library in order to read gzip compressed XML files. More...
|
| |
| class | HasActivationMethod |
| | Predicate that determines if a spectrum was generated using any activation method given in the constructor list. More...
|
| |
| class | HashGrid |
| | Container for (2-dimensional coordinate, value) pairs. More...
|
| |
| class | HasMetaValue |
| | Predicate that determines if a class has a certain metavalue. More...
|
| |
| class | HasPrecursorCharge |
| | Predicate that determines if a spectrum has a certain precursor charge as given in the constructor list. More...
|
| |
| class | HasScanMode |
| | Predicate that determines if a spectrum has a certain scan mode. More...
|
| |
| class | HasScanPolarity |
| | Predicate that determines if a spectrum has a certain scan polarity. More...
|
| |
| class | HDF5Connector |
| | File adapter for HDF5 files. More...
|
| |
| class | HiddenMarkovModel |
| | Hidden Markov Model implementation of PILIS. More...
|
| |
| class | HistogramDialog |
| | Dialog that show a HistogramWidget. More...
|
| |
| class | HistogramWidget |
| | Widget which can visualize a histogram. More...
|
| |
| class | HMMState |
| | Hidden Markov Model State class for the Hidden Markov Model. More...
|
| |
| class | HPLC |
| | Representation of a HPLC experiment. More...
|
| |
| class | HPLCVisualizer |
| | Class that displays all meta information for HPLC objects. More...
|
| |
| struct | HyperScore |
| | An implementation of the X!Tandem HyperScore PSM scoring function. More...
|
| |
| class | IBSpectraFile |
| | Implements the export of consensusmaps into the IBSpectra format used by isobar to load quantification results. More...
|
| |
| class | ICPLLabeler |
| | Simulate ICPL experiments. More...
|
| |
| class | IDConflictResolverAlgorithm |
| |
| class | IDDecoyProbability |
| | IDDecoyProbability calculates probabilities using decoy approach. More...
|
| |
| class | Identification |
| | Represents a object which can store the information of an analysisXML instance. More...
|
| |
| class | IdentificationData |
| | Representation of spectrum identification results and associated data. More...
|
| |
| class | IdentificationDataConverter |
| |
| class | IdentificationHit |
| | Represents a object which can store the information of an analysisXML instance. More...
|
| |
| class | IDFilter |
| | Collection of functions for filtering peptide and protein identifications. More...
|
| |
| class | IDMapper |
| | Annotates an MSExperiment, FeatureMap or ConsensusMap with peptide identifications. More...
|
| |
| class | IDMergerAlgorithm |
| | Creates a new Protein ID run into which other runs can be inserted. Creates union of protein hits but concatenates PSMs. Checks search engine consistency of all inserted runs. It differs from the IDMerger tool, in that it is an algorithm class and it allows inserting multiple peptide hits per peptide sequence (not only the first occurrence). More...
|
| |
| class | IDRipper |
| | Ripping protein/peptide identification according their file origin. More...
|
| |
| class | IDScoreGetterSetter |
| | A class for extracting and reinserting IDScores from Peptide/ProteinIdentifications and from ConsensusMaps. More...
|
| |
| class | IdXMLFile |
| | Used to load and store idXML files. More...
|
| |
| class | ILPDCWrapper |
| |
| class | IncludeExcludeTarget |
| | This class stores a SRM/MRM transition. More...
|
| |
| class | InclusionExclusionList |
| | Provides functionality for writing inclusion or exclusion lists. More...
|
| |
| class | IndexedMzMLDecoder |
| | A class to analyze indexedmzML files and extract the offsets of individual tags. More...
|
| |
| class | IndexedMzMLFileLoader |
| | A class to load an indexedmzML file. More...
|
| |
| class | INIFileEditorWindow |
| | shows the ParamEditor widget in a QMainWindow with a toolbar More...
|
| |
| class | InIntensityRange |
| | Predicate that determines if a peak lies inside/outside a specific intensity range. More...
|
| |
| class | INIUpdater |
| |
| class | InMSLevelRange |
| | Predicate that determines if a spectrum lies inside/outside a specific MS level set. More...
|
| |
| class | InMzRange |
| | Predicate that determines if a peak lies inside/outside a specific m/z range. More...
|
| |
| class | InPrecursorMZRange |
| | Predicate that determines if a spectrum's precursor is within a certain m/z range. More...
|
| |
| class | InRTRange |
| | Predicate that determines if a spectrum lies inside/outside a specific retention time range. More...
|
| |
| class | InspectInfile |
| | Inspect input file adapter. More...
|
| |
| class | InspectOutfile |
| | Representation of an Inspect outfile. More...
|
| |
| class | Instrument |
| | Description of a MS instrument. More...
|
| |
| class | InstrumentSettings |
| | Description of the settings a MS Instrument was run with. More...
|
| |
| class | InstrumentSettingsVisualizer |
| | Class that displays all meta information for InstrumentSettings objects. More...
|
| |
| class | InstrumentVisualizer |
| | Class that displays all meta information for an MS instrument. More...
|
| |
| class | IntensityBalanceFilter |
| | IntensityBalanceFilter divides the m/z-range into ten regions and sums the intensity in these regions. More...
|
| |
| class | InternalCalibration |
| | A mass recalibration method using linear/quadratic interpolation (robust/weighted) of given reference masses. More...
|
| |
| class | InterpolationModel |
| | Abstract class for 1D-models that are approximated using linear interpolation. More...
|
| |
| class | IonDetector |
| | Description of a ion detector (part of a MS Instrument) More...
|
| |
| class | IonDetectorVisualizer |
| | Class that displays all meta information for IonDetector objects. More...
|
| |
| class | IonizationSimulation |
| | Simulates Protein ionization. More...
|
| |
| class | IonMobilityScoring |
| | A class that calls the ion mobility scoring routines. More...
|
| |
| class | IonSource |
| | Description of an ion source (part of a MS Instrument) More...
|
| |
| class | IonSourceVisualizer |
| | Class that displays all meta information for IonSource objects. More...
|
| |
| class | IsEmptySpectrum |
| | Predicate that determines if a spectrum is empty. More...
|
| |
| class | IsInCollisionEnergyRange |
| | Predicate that determines if an MSn spectrum was generated with a collision energy in the given range. More...
|
| |
| class | IsInIsolationWindow |
| | Predicate that determines if the isolation window covers ANY of the given m/z values. More...
|
| |
| class | IsInIsolationWindowSizeRange |
| | Predicate that determines if the width of the isolation window of an MSn spectrum is in the given range. More...
|
| |
| class | IsobaricChannelExtractor |
| | Extracts individual channels from MS/MS spectra for isobaric labeling experiments. More...
|
| |
| class | IsobaricIsotopeCorrector |
| | Performs isotope impurity correction on the intensities extracted from an isobaric labeling experiment. More...
|
| |
| class | IsobaricNormalizer |
| | Performs median normalization on the extracted ratios of isobaric labeling experiment. More...
|
| |
| class | IsobaricQuantifier |
| | Given the extracted channel intensities the IsobaricQuantifier corrects and normalizes the intensities for further processing. More...
|
| |
| struct | IsobaricQuantifierStatistics |
| | Statistics for quantitation performance and comparison of NNLS vs. naive method (aka matrix inversion) More...
|
| |
| class | IsobaricQuantitationMethod |
| | Abstract base class describing an isobaric quantitation method in terms of the used channels and an isotope correction matrix. More...
|
| |
| class | IsoSpecGeneratorWrapper |
| | Interface for the IsoSpec algorithm - a generator of infinitely-resolved theoretical spectra. More...
|
| |
| class | IsoSpecOrderedGeneratorWrapper |
| | Generate the stream of configurations, ordered from most likely to least likely. More...
|
| |
| class | IsoSpecThresholdGeneratorWrapper |
| | Provides a threshold-based generator of isotopologues: generates all isotopologues more probable than a given probability threshold. More...
|
| |
| class | IsoSpecThresholdWrapper |
| | A non-generator version of IsoSpecThresholdGeneratorWrapper. More...
|
| |
| class | IsoSpecTotalProbGeneratorWrapper |
| | Generate a p-set of configurations for a given p (that is, a set of configurations such that their probabilities sum up to p). The p in normal usage will usually be close to 1 (e.g. 0.99). More...
|
| |
| class | IsoSpecTotalProbWrapper |
| | Create a p-set of configurations for a given p (that is, a set of configurations such that their probabilities sum up to p). The p in normal usage will usually be close to 1 (e.g. 0.99). More...
|
| |
| class | IsoSpecWrapper |
| | A convenience class for the IsoSpec algorithm - easier to use than the IsoSpecGeneratorWrapper classes. More...
|
| |
| struct | IsotopeCluster |
| | Stores information about an isotopic cluster (i.e. potential peptide charge variants) More...
|
| |
| class | IsotopeDiffFilter |
| | IsotopeDiffFilter returns total intensity of peak pairs that could result from isotope peaks. More...
|
| |
| class | IsotopeDistribution |
| |
| class | IsotopeDistributionCache |
| | Pre-calculate isotope distributions for interesting mass ranges. More...
|
| |
| class | IsotopeFitter1D |
| | Isotope distribution fitter (1-dim.) approximated using linear interpolation. More...
|
| |
| class | IsotopeMarker |
| | IsotopeMarker marks peak pairs which could represent an ion and its isotope. More...
|
| |
| class | IsotopeModel |
| | Isotope distribution approximated using linear interpolation. More...
|
| |
| class | IsotopePatternGenerator |
| | Provides an interface for different isotope pattern generator methods. More...
|
| |
| class | IsotopeWavelet |
| | Implements the isotope wavelet function. More...
|
| |
| class | IsotopeWaveletTransform |
| | A class implementing the isotope wavelet transform. If you just want to find features using the isotope wavelet, take a look at the FeatureFinderAlgorithmIsotopeWavelet class. Usually, you only have to consider the class at hand if you plan to change the basic implementation of the transform. More...
|
| |
| class | IsotopicDist |
| |
| class | IsZoomSpectrum |
| | Predicate that determines if a spectrum is a zoom (enhanced resolution) spectrum. More...
|
| |
| class | ItraqConstants |
| | Some constants used throughout iTRAQ classes. More...
|
| |
| class | ItraqEightPlexQuantitationMethod |
| | iTRAQ 8 plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | ItraqFourPlexQuantitationMethod |
| | iTRAQ 4 plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | ITRAQLabeler |
| | Simulate iTRAQ experiments. More...
|
| |
| class | JavaInfo |
| | Detect Java and retrieve information. More...
|
| |
| class | KDTreeFeatureMaps |
| | Stores a set of features, together with a 2D tree for fast search. More...
|
| |
| class | KDTreeFeatureNode |
| | A node of the kD-tree with pointer to corresponding data and index. More...
|
| |
| class | KroenikFile |
| | File adapter for Kroenik (HardKloer sibling) files. More...
|
| |
| class | LabeledPairFinder |
| | The LabeledPairFinder allows the matching of labeled features (features with a fixed distance). More...
|
| |
| class | LabelFreeLabeler |
| | Abstract base class for all kinds of labeling techniques. More...
|
| |
| class | LayerData |
| | Class that stores the data for one layer. More...
|
| |
| class | LayerStatisticsDialog |
| | Dialog showing statistics about the data of the current layer. More...
|
| |
| class | LCElutionPeak |
| |
| class | LCMS |
| |
| class | LCMSCData |
| |
| class | LevMarqFitter1D |
| | Abstract class for 1D-model fitter using Levenberg-Marquardt algorithm for parameter optimization. More...
|
| |
| struct | LexicographicComparator |
| | A wrapper class that combines two comparators lexicographically. Normally you should use the make-function lexicographicComparator() because then you do not need to specify the template arguments. More...
|
| |
| class | LibSVMEncoder |
| | Serves for encoding sequences into feature vectors. More...
|
| |
| class | LinearResampler |
| | Linear Resampling of raw data. More...
|
| |
| class | LinearResamplerAlign |
| | Linear Resampling of raw data with alignment. More...
|
| |
| class | ListEditor |
| | Editor for editing int, double and string lists (including output and input file lists) More...
|
| |
| class | ListUtils |
| | Collection of utility functions for management of vectors. More...
|
| |
| class | LocalLinearMap |
| | Trained Local Linear Map (LLM) model for peak intensity prediction. More...
|
| |
| class | LogConfigHandler |
| | The LogConfigHandler provides the functionality to configure the internal logging of OpenMS algorithms that use the global instances of LogStream. More...
|
| |
| class | LowessSmoothing |
| | LOWESS (locally weighted scatterplot smoothing). More...
|
| |
| class | LPWrapper |
| |
| class | Map |
| | Map class based on the STL map (containing several convenience functions) More...
|
| |
| class | MapAlignmentAlgorithmIdentification |
| | A map alignment algorithm based on peptide identifications from MS2 spectra. More...
|
| |
| class | MapAlignmentAlgorithmKD |
| | An efficient reference-free feature map alignment algorithm for unlabeled data. More...
|
| |
| class | MapAlignmentAlgorithmPoseClustering |
| | A map alignment algorithm based on pose clustering. More...
|
| |
| class | MapAlignmentAlgorithmSpectrumAlignment |
| | A map alignment algorithm based on spectrum similarity (dynamic programming). More...
|
| |
| class | MapAlignmentAlgorithmTreeGuided |
| | A map alignment algorithm based on peptide identifications from MS2 spectra. More...
|
| |
| class | MapAlignmentEvaluationAlgorithm |
| | Base class for all Caap evaluation algorithms. More...
|
| |
| class | MapAlignmentEvaluationAlgorithmPrecision |
| | Caap evaluation algorithm to obtain a precision value. More...
|
| |
| class | MapAlignmentEvaluationAlgorithmRecall |
| | Caap evaluation algorithm to obtain a recall value. More...
|
| |
| class | MapAlignmentTransformer |
| | This class collects functions for applying retention time transformations to data structures. More...
|
| |
| class | MapConversion |
| |
| class | MarkerMower |
| | MarkerMower uses PeakMarker to find peaks, those that are not marked get removed. More...
|
| |
| class | MascotGenericFile |
| | Read/write Mascot generic files (MGF). More...
|
| |
| class | MascotInfile |
| | Mascot input file adapter. More...
|
| |
| class | MascotRemoteQuery |
| | Class which handles the communication between OpenMS and the Mascot server. More...
|
| |
| class | MascotXMLFile |
| | Used to load Mascot XML files. More...
|
| |
| class | MassAnalyzer |
| | Description of a mass analyzer (part of a MS Instrument) More...
|
| |
| class | MassAnalyzerVisualizer |
| | Class that displays all meta information for MassAnalyzer objects. More...
|
| |
| class | MassDecomposition |
| | Class represents a decomposition of a mass into amino acids. More...
|
| |
| class | MassDecompositionAlgorithm |
| | Mass decomposition algorithm, given a mass it suggests possible compositions. More...
|
| |
| class | MassExplainer |
| | computes empirical formulas for given mass differences using a set of allowed elements More...
|
| |
| class | MassTrace |
| | A container type that gathers peaks similar in m/z and moving along retention time. More...
|
| |
| class | MasstraceCorrelator |
| | Correlates individual masstraces found in mass spectrometric maps. More...
|
| |
| class | MassTraceDetection |
| | A mass trace extraction method that gathers peaks similar in m/z and moving along retention time. More...
|
| |
| class | MatchedIterator |
| | For each element in the reference container the closest peak in the target will be searched. If no match is found within the tolerance window, the peak will be skipped over. More...
|
| |
| class | Matrix |
| | A two-dimensional matrix. Similar to std::vector, but uses a binary operator(,) for element access. More...
|
| |
| class | MaxLikeliFitter1D |
| | Abstract base class for all 1D-model fitters using maximum likelihood optimization. More...
|
| |
| class | MetaboliteFeatureDeconvolution |
| | An algorithm to decharge small molecule features (i.e. as found by FeatureFinder). More...
|
| |
| class | MetaboliteSpectralMatching |
| |
| class | MetaboTargetedAssay |
| | This class provides methods for the extraction of targeted assays for metabolomics. More...
|
| |
| class | MetaDataBrowser |
| | A meta data visualization widget. More...
|
| |
| class | MetaInfo |
| | A Type-Name-Value tuple class. More...
|
| |
| class | MetaInfoDescription |
| | Description of the meta data arrays of MSSpectrum. More...
|
| |
| class | MetaInfoDescriptionVisualizer |
| | Class that displays all meta information for MetaInfoDescription objects. More...
|
| |
| class | MetaInfoInterface |
| | Interface for classes that can store arbitrary meta information (Type-Name-Value tuples). More...
|
| |
| class | MetaInfoInterfaceUtils |
| | Utilities operating on containers inheriting from MetaInfoInterface. More...
|
| |
| class | MetaInfoRegistry |
| | Registry which assigns unique integer indices to strings. More...
|
| |
| class | MetaInfoVisualizer |
| | MetaInfoVisualizer is a visualizer class for all classes that use one MetaInfo object as member. More...
|
| |
| class | MinimumDistance |
| | basic data structure for distances between clusters More...
|
| |
| class | MissedCleavages |
| | This class is a metric for the QualityControl TOPP Tool. More...
|
| |
| class | ModelDescription |
| | Stores the name and parameters of a model. More...
|
| |
| class | Modification |
| | Meta information about chemical modification of a sample. More...
|
| |
| class | ModificationDefinition |
| | Representation of modification definition. More...
|
| |
| class | ModificationDefinitionsSet |
| | Representation of a set of modification definitions. More...
|
| |
| class | ModificationsDB |
| | database which holds all residue modifications from UniMod More...
|
| |
| class | ModificationVisualizer |
| | Class that displays all meta information of modification objects. More...
|
| |
| class | ModifiedNASequenceGenerator |
| |
| class | ModifiedPeptideGenerator |
| |
| struct | MorpheusScore |
| | An implementation of the Morpheus PSM scoring function Inspired by a C# implementation by C. Wenger released under MIT license. More...
|
| |
| class | MorphologicalFilter |
| | This class implements baseline filtering operations using methods from mathematical morphology. More...
|
| |
| class | MRMAssay |
| | Generate assays from a TargetedExperiment. More...
|
| |
| class | MRMBatchFeatureSelector |
| |
| class | MRMDecoy |
| | This class generates a TargetedExperiment object with decoys based on a TargetedExperiment object. More...
|
| |
| class | MRMFeature |
| | A multi-chromatogram MRM feature. More...
|
| |
| class | MRMFeatureFilter |
| | The MRMFeatureFilter either flags components and/or transitions that do not pass the QC criteria or filters out components and/or transitions that do not pass the QC criteria. More...
|
| |
| class | MRMFeatureFinderScoring |
| | The MRMFeatureFinder finds and scores peaks of transitions that co-elute. More...
|
| |
| class | MRMFeatureOpenMS |
| | An implementation of the OpenSWATH MRM Feature Access interface using OpenMS. More...
|
| |
| class | MRMFeaturePicker |
| | _MRMFeaturePicker_ defines the structures containing parameters to be used in [MRMTransitionGroupPicker](MRMTransitionGroupPicker) for components and components groups. More...
|
| |
| class | MRMFeaturePickerFile |
| | _MRMFeaturePickerFile_ loads components and components groups parameters from a .csv file. More...
|
| |
| class | MRMFeatureQC |
| | The MRMFeatureQC is a class to handle the parameters and options for MRMFeatureFilter. More...
|
| |
| class | MRMFeatureQCFile |
| | File adapter for MRMFeatureQC files. More...
|
| |
| class | MRMFeatureSelector |
| |
| class | MRMFeatureSelector_test |
| |
| class | MRMFeatureSelectorQMIP |
| |
| class | MRMFeatureSelectorScore |
| |
| class | MRMFragmentSelection |
| | This class can select appropriate fragment ions of an MS/MS spectrum of a peptide. More...
|
| |
| class | MRMIonSeries |
| | Generate theoretical fragment ion series for use in MRMAssay and MRMDecoy. More...
|
| |
| class | MRMMapping |
| | A class to map targeted assays to chromatograms. More...
|
| |
| class | MRMRTNormalizer |
| | The MRMRTNormalizer will find retention time peptides in data. More...
|
| |
| class | MRMTransitionGroup |
| | The representation of a group of transitions in a targeted proteomics experiment. More...
|
| |
| class | MRMTransitionGroupPicker |
| | The MRMTransitionGroupPicker finds peaks in chromatograms that belong to the same precursors. More...
|
| |
| class | MS1FeatureMerger |
| |
| struct | MS1Signal |
| |
| class | MS2ConsensusSpectrum |
| |
| class | MS2Feature |
| |
| class | MS2File |
| | MS2 input file adapter. More...
|
| |
| class | MS2Fragment |
| |
| class | Ms2IdentificationRate |
| | This class is a metric for the QualityControl-ToppTool. More...
|
| |
| class | MS2Info |
| |
| class | Ms2SpectrumStats |
| | QC metric to determine the number of MS2 scans per MS1 scan over RT. More...
|
| |
| class | MSChromatogram |
| | The representation of a chromatogram. More...
|
| |
| class | MSDataAggregatingConsumer |
| | Aggregates spectra by retention time. More...
|
| |
| class | MSDataCachedConsumer |
| | Transforming and cached writing consumer of MS data. More...
|
| |
| class | MSDataChainingConsumer |
| | Consumer class that passes all consumed data through a set of operations. More...
|
| |
| class | MSDataSqlConsumer |
| | A data consumer that inserts MS data into a SQLite database. More...
|
| |
| class | MSDataStoringConsumer |
| | Consumer class that simply stores the data. More...
|
| |
| class | MSDataTransformingConsumer |
| | Transforming consumer of MS data. More...
|
| |
| class | MSDataWritingConsumer |
| | Consumer class that writes MS data to disk using the mzML format. More...
|
| |
| class | MSExperiment |
| | In-Memory representation of a mass spectrometry experiment. More...
|
| |
| class | MsInspectFile |
| | File adapter for MsInspect files. More...
|
| |
| class | MSNumpressCoder |
| | Class to encode and decode data encoded with MSNumpress. More...
|
| |
| class | MSPeak |
| |
| class | MSPFile |
| | File adapter for MSP files (NIST spectra library) More...
|
| |
| class | MSPGenericFile |
| | Load MSP text file and save it into an `MSExperiment`. More...
|
| |
| class | MSPGenericFile_friend |
| |
| class | MSQuantifications |
| |
| class | MSSim |
| | Central class for simulation of mass spectrometry experiments. More...
|
| |
| class | MSSpectrum |
| | The representation of a 1D spectrum. More...
|
| |
| class | MSstatsFile |
| | File adapter for MzTab files. More...
|
| |
| class | MultiGradient |
| | A gradient of multiple colors and arbitrary distances between colors. More...
|
| |
| class | MultiGradientSelector |
| | A widget witch allows constructing gradients of multiple colors. More...
|
| |
| class | MultiplexClustering |
| | clusters results from multiplex filtering More...
|
| |
| class | MultiplexDeltaMasses |
| | data structure for mass shift pattern More...
|
| |
| class | MultiplexDeltaMassesGenerator |
| | generates complete list of all possible mass shifts due to isotopic labelling More...
|
| |
| class | MultiplexFilteredMSExperiment |
| | data structure storing all peaks (and optionally their raw data points) of an experiment corresponding to one specific peak pattern More...
|
| |
| class | MultiplexFilteredPeak |
| | data structure storing a single peak that passed all filters More...
|
| |
| class | MultiplexFiltering |
| | base class for filtering centroided and profile data for peak patterns More...
|
| |
| class | MultiplexFilteringCentroided |
| | filters centroided data for peak patterns More...
|
| |
| class | MultiplexFilteringProfile |
| | filters centroided and profile data for peak patterns More...
|
| |
| class | MultiplexIsotopicPeakPattern |
| | data structure for pattern of isotopic peaks More...
|
| |
| class | MultiplexSatelliteCentroided |
| | data structure storing a single satellite peak More...
|
| |
| class | MultiplexSatelliteProfile |
| | data structure storing a single satellite data point More...
|
| |
| class | MzCalibration |
| | QC metric calculating (un)calibrated m/z error. More...
|
| |
| class | MzDataFile |
| | File adapter for MzData files. More...
|
| |
| class | MzIdentMLFile |
| | File adapter for MzIdentML files. More...
|
| |
| class | MzMLFile |
| | File adapter for MzML files. More...
|
| |
| class | MzMLSpectrumDecoder |
| | A class to decode input strings that contain an mzML chromatogram or spectrum tag. More...
|
| |
| class | MzMLSwathFileConsumer |
| | On-disk mzML implementation of FullSwathFileConsumer. More...
|
| |
| class | MzQuantMLFile |
| | File adapter for MzQuantML files. More...
|
| |
| class | MzTab |
| | Data model of MzTab files. Please see the official MzTab specification at https://code.google.com/p/mztab/. More...
|
| |
| struct | MzTabAssayMetaData |
| |
| class | MzTabBoolean |
| |
| struct | MzTabContactMetaData |
| |
| struct | MzTabCVMetaData |
| |
| class | MzTabDouble |
| |
| class | MzTabDoubleList |
| |
| class | MzTabFile |
| | File adapter for MzTab files. More...
|
| |
| struct | MzTabInstrumentMetaData |
| |
| class | MzTabInteger |
| |
| class | MzTabIntegerList |
| |
| class | MzTabMetaData |
| | all meta data of a mzTab file. Please refer to specification for documentation. More...
|
| |
| class | MzTabModification |
| |
| class | MzTabModificationList |
| |
| struct | MzTabModificationMetaData |
| |
| struct | MzTabMSRunMetaData |
| |
| struct | MzTabNucleicAcidSectionRow |
| | NUC - Nucleic acid section (table-based) More...
|
| |
| class | MzTabNullAbleBase |
| | base class for atomic, non-container types (Double, Int) More...
|
| |
| class | MzTabNullAbleInterface |
| | basic interface for all MzTab datatypes (can be null; are converted from and to cell string) More...
|
| |
| class | MzTabNullNaNAndInfAbleBase |
| | base class for the atomic non-container like MzTab data types (Double, Int) More...
|
| |
| class | MzTabNullNaNAndInfAbleInterface |
| | interface for NaN- and Inf- able datatypes (Double and Integer in MzTab). These are as well null-able More...
|
| |
| struct | MzTabOligonucleotideSectionRow |
| | OLI - Oligonucleotide section (table-based) More...
|
| |
| struct | MzTabOSMSectionRow |
| | OSM - OSM (oligonucleotide-spectrum match) section (table-based) More...
|
| |
| class | MzTabParameter |
| |
| class | MzTabParameterList |
| |
| struct | MzTabPeptideSectionRow |
| | PEP - Peptide section (Table based) More...
|
| |
| struct | MzTabProteinSectionRow |
| | PRT - Protein section (Table based) More...
|
| |
| struct | MzTabPSMSectionRow |
| | PSM - PSM section (Table based) More...
|
| |
| struct | MzTabSampleMetaData |
| |
| struct | MzTabSmallMoleculeSectionRow |
| | SML Small molecule section (table based) More...
|
| |
| struct | MzTabSoftwareMetaData |
| |
| class | MzTabSpectraRef |
| |
| class | MzTabString |
| |
| class | MzTabStringList |
| |
| struct | MzTabStudyVariableMetaData |
| |
| class | MZTrafoModel |
| | Create and apply models of a mass recalibration function. More...
|
| |
| class | MzXMLFile |
| | File adapter for MzXML 3.1 files. More...
|
| |
| class | NASequence |
| | Representation of a nucleic acid sequence. More...
|
| |
| class | NetworkGetRequest |
| |
| class | NeutralLossDiffFilter |
| | NeutralLossDiffFilter returns the total intensity ob peak pairs whose m/z difference can be explained by a neutral loss. More...
|
| |
| class | NeutralLossMarker |
| | NeutralLossMarker marks peak pairs which could represent an ion an its neutral loss (water, ammonia) More...
|
| |
| class | NLargest |
| | NLargest removes all but the n largest peaks. More...
|
| |
| class | NonNegativeLeastSquaresSolver |
| | Wrapper for a non-negative least squares (NNLS) solver. More...
|
| |
| class | NoopMSDataConsumer |
| | Consumer class that performs no operation. More...
|
| |
| class | NoopMSDataWritingConsumer |
| | Consumer class that perform no operation. More...
|
| |
| class | Normalizer |
| | Normalizes the peak intensities spectrum-wise. More...
|
| |
| class | NucleicAcidSpectrumGenerator |
| | Generates theoretical spectra for nucleic acid sequences. More...
|
| |
| class | O18Labeler |
| | Simulate O-18 experiments. More...
|
| |
| class | OfflinePrecursorIonSelection |
| | Implements different algorithms for precursor ion selection. More...
|
| |
| class | OMSSACSVFile |
| | File adapter for OMSSACSV files. More...
|
| |
| class | OMSSAXMLFile |
| | Used to load OMSSAXML files. More...
|
| |
| class | OnDiscMSExperiment |
| | Representation of a mass spectrometry experiment on disk. More...
|
| |
| class | OpenPepXLAlgorithm |
| |
| class | OpenPepXLLFAlgorithm |
| |
| struct | OpenSwath_Ind_Scores |
| |
| struct | OpenSwath_Scores |
| | A structure to hold the different scores computed by OpenSWATH. More...
|
| |
| struct | OpenSwath_Scores_Usage |
| | A structure to store which scores should be used by the OpenSWATH Algorithm. More...
|
| |
| class | OpenSwathCalibrationWorkflow |
| | Execute all steps for retention time and m/z calibration of SWATH-MS data. More...
|
| |
| class | OpenSwathDataAccessHelper |
| | Several helpers to convert OpenMS datastructures to structures that implement the OpenSWATH interfaces. More...
|
| |
| class | OpenSwathHelper |
| | A helper class that is used by several OpenSWATH tools. More...
|
| |
| class | OpenSwathOSWWriter |
| | Class to write out an OpenSwath OSW SQLite output (PyProphet input). More...
|
| |
| class | OpenSwathScoring |
| | A class that calls the scoring routines. More...
|
| |
| class | OpenSwathTSVWriter |
| | Class to write out an OpenSwath TSV output (mProphet input). More...
|
| |
| class | OpenSwathWorkflow |
| | Execute all steps in an OpenSwath analysis. More...
|
| |
| class | OpenSwathWorkflowBase |
| |
| class | OpenSwathWorkflowSonar |
| | Execute all steps in an OpenEcho analysis (OpenSwath for SONAR data) More...
|
| |
| class | OptimizePeakDeconvolution |
| | This class provides the deconvolution of peak regions using non-linear optimization. More...
|
| |
| class | OptimizePick |
| | This class provides the non-linear optimization of the peak parameters. More...
|
| |
| class | OPXLDataStructs |
| |
| class | OPXLHelper |
| | The OPXLHelper class contains functions needed by OpenPepXL and OpenPepXLLF to reduce duplicated code. More...
|
| |
| class | OPXLSpectrumProcessingAlgorithms |
| |
| class | OSWFile |
| | This class serves for reading in and writing OpenSWATH OSW files. More...
|
| |
| struct | PairComparatorFirstElement |
| | Class for comparison of std::pair using first ONLY e.g. for use with std::sort. More...
|
| |
| struct | PairComparatorFirstElementMore |
| | Class for comparison of std::pair using first ONLY e.g. for use with std::sort. More...
|
| |
| struct | PairComparatorSecondElement |
| | Class for comparison of std::pair using second ONLY e.g. for use with std::sort. More...
|
| |
| struct | PairComparatorSecondElementMore |
| | Class for comparison of std::pair using second ONLY e.g. for use with std::sort. More...
|
| |
| struct | PairMatcherFirstElement |
| | Class for comparison of std::pair using first ONLY e.g. for use with std::sort. More...
|
| |
| struct | PairMatcherSecondElement |
| | Struct for comparison of std::pair using second ONLY e.g. for use with std::sort. More...
|
| |
| class | Param |
| | Management and storage of parameters / INI files. More...
|
| |
| class | ParamEditor |
| | A GUI for editing or viewing a Param object. More...
|
| |
| struct | ParameterInformation |
| | Struct that captures all information of a command line parameter. More...
|
| |
| class | ParamXMLFile |
| | The file pendant of the Param class used to load and store the param datastructure as paramXML. More...
|
| |
| class | ParentPeakMower |
| | ParentPeakMower gets rid of high peaks that could stem from unfragmented precursor ions. More...
|
| |
| class | Peak1D |
| | A 1-dimensional raw data point or peak. More...
|
| |
| class | Peak2D |
| | A 2-dimensional raw data point or peak. More...
|
| |
| class | PeakAlignment |
| | make a PeakAlignment of two PeakSpectra More...
|
| |
| struct | PeakCandidate |
| | A small structure to hold peak candidates. More...
|
| |
| class | PeakFileOptions |
| | Options for loading files containing peak data. More...
|
| |
| struct | PeakIndex |
| | Index of a peak or feature. More...
|
| |
| class | PeakIntegrator |
| | Compute the area, background and shape metrics of a peak. More...
|
| |
| class | PeakIntensityPredictor |
| | Predict peak heights of peptides based on Local Linear Map model. More...
|
| |
| class | PeakMarker |
| | PeakMarker marks peaks that seem to fulfill some criterion. More...
|
| |
| class | PeakPickerCWT |
| | This class implements a peak picking algorithm using wavelet techniques. More...
|
| |
| class | PeakPickerHiRes |
| | This class implements a fast peak-picking algorithm best suited for high resolution MS data (FT-ICR-MS, Orbitrap). In high resolution data, the signals of ions with similar mass-to-charge ratios (m/z) exhibit little or no overlapping and therefore allow for a clear separation. Furthermore, ion signals tend to show well-defined peak shapes with narrow peak width. More...
|
| |
| class | PeakPickerIterative |
| | This class implements a peak-picking algorithm for high-resolution MS data (specifically designed for TOF-MS data). More...
|
| |
| class | PeakPickerMaxima |
| | This class implements a fast peak-picking algorithm best suited for high resolution MS data (FT-ICR-MS, Orbitrap). In high resolution data, the signals of ions with similar mass-to-charge ratios (m/z) exhibit little or no overlapping and therefore allow for a clear separation. Furthermore, ion signals tend to show well-defined peak shapes with narrow peak width. More...
|
| |
| class | PeakPickerMRM |
| | The PeakPickerMRM finds peaks a single chromatogram. More...
|
| |
| class | PeakPickerSH |
| |
| class | PeakShape |
| | Internal representation of a peak shape (used by the PeakPickerCWT) More...
|
| |
| class | PeakSpectrumCompareFunctor |
| | Base class for compare functors of spectra, that return a similarity value for two spectra. More...
|
| |
| class | PeakTypeEstimator |
| | Estimates if the data of a spectrum is raw data or peak data. More...
|
| |
| class | PeakWidthEstimator |
| | Rough estimation of the peak width at m/z. More...
|
| |
| class | PepNovoInfile |
| | PepNovo input file adapter. More...
|
| |
| class | PepNovoOutfile |
| | Representation of a PepNovo output file. More...
|
| |
| class | PeptideAndProteinQuant |
| | Helper class for peptide and protein quantification based on feature data annotated with IDs. More...
|
| |
| class | PeptideEvidence |
| | Representation of a peptide evidence. More...
|
| |
| class | PeptideHit |
| | Representation of a peptide hit. More...
|
| |
| class | PeptideHitVisualizer |
| | Class that displays all meta information for PeptideHit objects. More...
|
| |
| class | PeptideIdentification |
| | Represents the peptide hits for a spectrum. More...
|
| |
| class | PeptideIdentificationVisualizer |
| | Class that displays all meta information for PeptideIdentification objects. More...
|
| |
| class | PeptideIndexing |
| | Refreshes the protein references for all peptide hits in a vector of PeptideIdentifications and adds target/decoy information. More...
|
| |
| class | PeptideProteinResolution |
| | Resolves shared peptides based on protein scores. More...
|
| |
| class | PepXMLFile |
| | Used to load and store PepXML files. More...
|
| |
| class | PepXMLFileMascot |
| | Used to load Mascot PepXML files. More...
|
| |
| class | PercolatorFeatureSetHelper |
| | Percolator feature set and integration helper. More...
|
| |
| class | PercolatorOutfile |
| | Class for reading Percolator tab-delimited output files. More...
|
| |
| class | PlainMSDataWritingConsumer |
| | Consumer class that writes MS data to disk using the mzML format. More...
|
| |
| struct | PointerComparator |
| | Wrapper that takes a comparator for `something' and makes a comparator for pointers to `something' out of it. Normally you should use the make-function pointerComparator() because then you do not need to specify the template arguments. More...
|
| |
| class | PoseClusteringAffineSuperimposer |
| | A superimposer that uses a voting scheme, also known as pose clustering, to find a good affine transformation. More...
|
| |
| class | PoseClusteringShiftSuperimposer |
| | A superimposer that uses a voting scheme, also known as pose clustering, to find a good shift transformation. More...
|
| |
| struct | PpmTrait |
| |
| struct | PrecisionWrapper |
| | Wrapper class to implement output with appropriate precision. See precisionWrapper(). More...
|
| |
| class | Precursor |
| | Precursor meta information. More...
|
| |
| class | PrecursorCorrection |
| | This class provides methods for precursor correction. More...
|
| |
| class | PrecursorIonSelection |
| | This class implements different precursor ion selection strategies. More...
|
| |
| class | PrecursorIonSelectionPreprocessing |
| | This class implements the database preprocessing needing for precursor ion selection. More...
|
| |
| struct | PrecursorMassComparator |
| |
| class | PrecursorPurity |
| | Precursor purity or noise estimation. More...
|
| |
| class | PrecursorVisualizer |
| | Class that displays all meta information for Precursor objects. More...
|
| |
| struct | ProbablePhosphoSites |
| |
| class | ProcessData |
| |
| class | Product |
| | Product meta information. More...
|
| |
| class | ProductModel |
| | Class for product models i.e. models with D independent dimensions. More...
|
| |
| class | ProductModel< 2 > |
| | The class template is only implemented for D=2 because we use Peak2D here. More...
|
| |
| class | ProductVisualizer |
| | Class that displays all meta information for Product objects. More...
|
| |
| class | ProgressLogger |
| | Base class for all classes that want to report their progress. More...
|
| |
| class | ProteaseDB |
| | Database for enzymes that digest proteins (proteases) More...
|
| |
| class | ProteaseDigestion |
| | Class for the enzymatic digestion of proteins. More...
|
| |
| class | ProteinHit |
| | Representation of a protein hit. More...
|
| |
| class | ProteinHitVisualizer |
| | Class that displays all meta information for ProteinHit objects. More...
|
| |
| class | ProteinIdentification |
| | Representation of a protein identification run. More...
|
| |
| class | ProteinIdentificationVisualizer |
| | Class that displays all meta information for ProteinIdentification objects. More...
|
| |
| class | ProteinInference |
| | [experimental class] given a peptide quantitation, infer corresponding protein quantities More...
|
| |
| class | ProteinResolver |
| | Helper class for peptide and protein quantification based on feature data annotated with IDs. More...
|
| |
| class | ProtonDistributionModel |
| | A proton distribution model to calculate the proton distribution over charged peptides. More...
|
| |
| class | ProtXMLFile |
| | Used to load (storing not supported, yet) ProtXML files. More...
|
| |
| struct | PScore |
| | Implementation of the PScore PSM scoring algorithm. More...
|
| |
| class | PSLPFormulation |
| | Implements ILP formulation of precursor selection problems. More...
|
| |
| class | PSProteinInference |
| | This class implements protein inference for the precursor ion selection strategies. More...
|
| |
| class | PTMXMLFile |
| | Used to load and store PTMXML files. More...
|
| |
| class | QApplicationTOPP |
| | Extension to the QApplication for running TOPPs GUI tools. More...
|
| |
| class | QCBase |
| | This class serves as an abstract base class for all QC classes. More...
|
| |
| class | QcMLFile |
| | File adapter for QcML files used to load and store QcML files. More...
|
| |
| class | QTCluster |
| | A representation of a QT cluster used for feature grouping. More...
|
| |
| class | QTClusterFinder |
| | A variant of QT clustering for the detection of feature groups. More...
|
| |
| class | QuantitativeExperimentalDesign |
| | Merge files according to experimental design. More...
|
| |
| struct | Range |
| | Internal structure to store a lower and upper bound of an m/z range. More...
|
| |
| class | RangeManager |
| | Handles the management of a position and intensity range. More...
|
| |
| class | RawData |
| |
| class | RawMSSignalSimulation |
| | Simulates MS signals for a given set of peptides. More...
|
| |
| class | RawTandemMSSignalSimulation |
| | Simulates tandem MS signals for a given set of peptides. More...
|
| |
| class | ReactionMonitoringTransition |
| | This class stores a SRM/MRM transition. More...
|
| |
| class | RegularSwathFileConsumer |
| | In-memory implementation of FullSwathFileConsumer. More...
|
| |
| class | Residue |
| | Representation of a residue. More...
|
| |
| class | ResidueDB |
| | residue data base which holds residues More...
|
| |
| class | ResidueModification |
| | Representation of a modification. More...
|
| |
| struct | ReverseComparator |
| | Wrapper that reverses (exchanges) the two arguments of a comparator. Normally you should use the make-function reverseComparator() because then you do not need to specify the template arguments. More...
|
| |
| class | Ribonucleotide |
| | Representation of a ribonucleotide (modified or unmodified) More...
|
| |
| class | RibonucleotideDB |
| | Database of ribonucleotides (modified and unmodified) More...
|
| |
| class | RichPeak2D |
| | A 2-dimensional raw data point or peak with meta information. More...
|
| |
| class | RNaseDB |
| | Database for enzymes that digest RNA (RNases) More...
|
| |
| class | RNaseDigestion |
| | Class for the enzymatic digestion of RNAs. More...
|
| |
| struct | RNPxlMarkerIonExtractor |
| |
| struct | RNPxlModificationMassesResult |
| |
| class | RNPxlModificationsGenerator |
| |
| struct | RNPxlReport |
| | create report More...
|
| |
| struct | RNPxlReportRow |
| | struct to hold a single report line More...
|
| |
| struct | RNPxlReportRowHeader |
| | create header line More...
|
| |
| class | RTAlignment |
| | Take the original retention time before map alignment and use the alignment's trafoXML for calculation of the new alignment retention times. More...
|
| |
| class | RTSimulation |
| | Simulates/Predicts retention times for peptides or peptide separation. More...
|
| |
| class | RWrapper |
| | R-Wrapper Class. More...
|
| |
| class | Sample |
| | Meta information about the sample. More...
|
| |
| class | SampleTreatment |
| | Base class for sample treatments (Digestion, Modification, Tagging, ...) More...
|
| |
| class | SampleVisualizer |
| | Class that displays all meta information of sample objects. More...
|
| |
| class | SaveImageDialog |
| | Dialog for saving an image. More...
|
| |
| class | SavitzkyGolayFilter |
| | Computes the Savitzky-Golay filter coefficients using QR decomposition. More...
|
| |
| class | Scaler |
| | Scaler scales the peak by ranking the peaks and assigning intensity according to rank. More...
|
| |
| struct | ScanWindow |
| | Scan window description. More...
|
| |
| class | ScanWindowVisualizer |
| | Class that displays all meta information for ScanWindow objects. More...
|
| |
| struct | ScoreToTgtDecLabelPairs |
| |
| class | SeedListGenerator |
| | Generate seed lists for feature detection. More...
|
| |
| class | SequestInfile |
| | Sequest input file adapter. More...
|
| |
| class | SequestOutfile |
| | Representation of a Sequest output file. More...
|
| |
| class | SHFeature |
| |
| class | SignalToNoiseEstimator |
| | This class represents the abstract base class of a signal to noise estimator. More...
|
| |
| class | SignalToNoiseEstimatorMeanIterative |
| | Estimates the signal/noise (S/N) ratio of each data point in a scan based on an iterative scheme which discards high intensities. More...
|
| |
| class | SignalToNoiseEstimatorMedian |
| | Estimates the signal/noise (S/N) ratio of each data point in a scan by using the median (histogram based) More...
|
| |
| class | SignalToNoiseEstimatorMedianRapid |
| | Estimates the signal/noise (S/N) ratio of each data point in a scan by using the median (window based) More...
|
| |
| class | SignalToNoiseOpenMS |
| | An implementation of the OpenSWATH SignalToNoise Access interface using OpenMS. More...
|
| |
| class | SILACLabeler |
| | Simulate SILAC experiments. More...
|
| |
| class | SimpleOpenMSSpectraFactory |
| | A factory method that returns two ISpectrumAccess implementations. More...
|
| |
| class | SimplePairFinder |
| | This class implements a simple point pair finding algorithm. More...
|
| |
| class | SimpleSearchEngineAlgorithm |
| |
| class | SimpleSVM |
| | Simple interface to support vector machines for classification (via LIBSVM). More...
|
| |
| class | SimpleTSGXLMS |
| | Generates theoretical spectra for cross-linked peptides. More...
|
| |
| class | SingleLinkage |
| | SingleLinkage ClusterMethod. More...
|
| |
| class | SingletonRegistry |
| | Holds pointers to unique instance of a singleton factory. More...
|
| |
| class | SiriusAdapterAlgorithm |
| |
| class | SiriusFragmentAnnotation |
| |
| class | SiriusMSFile |
| |
| class | SiriusMzTabWriter |
| |
| class | Software |
| | Description of the software used for processing. More...
|
| |
| class | SoftwareVisualizer |
| | Class that displays all meta information for Software objects. More...
|
| |
| class | SONARScoring |
| | Scoring of an spectrum using SONAR data. More...
|
| |
| class | SourceFile |
| | Description of a file location, used to store the origin of (meta) data. More...
|
| |
| class | SourceFileVisualizer |
| | Class that displays all meta information for SourceFile objects. More...
|
| |
| class | SpecArrayFile |
| | File adapter for SpecArray (.pepList) files. More...
|
| |
| class | SpectraIdentificationViewWidget |
| | Tabular visualization / selection of identified spectra. More...
|
| |
| class | SpectralMatch |
| |
| struct | SpectralMatchScoreComparator |
| |
| class | SpectraMerger |
| | Merges blocks of MS or MS2 spectra. More...
|
| |
| class | SpectraSTSimilarityScore |
| | Similarity score of SpectraST. More...
|
| |
| class | SpectraViewWidget |
| | Hierarchical visualization and selection of spectra. More...
|
| |
| class | Spectrum1DCanvas |
| | Canvas for visualization of one or several spectra. More...
|
| |
| class | Spectrum1DGoToDialog |
| | simple goto/set visible area dialog for exact placement of the viewing window More...
|
| |
| class | Spectrum1DWidget |
| | Widget for visualization of several spectra. More...
|
| |
| class | Spectrum2DCanvas |
| | Canvas for 2D-visualization of peak map, feature map and consensus map data. More...
|
| |
| class | Spectrum2DGoToDialog |
| | GoTo dialog used to zoom to a m/z and retention time range or to a feature. More...
|
| |
| class | Spectrum2DWidget |
| | Widget for 2D-visualization of peak map and feature map data. More...
|
| |
| class | Spectrum3DCanvas |
| | Canvas for 3D-visualization of peak map data. More...
|
| |
| class | Spectrum3DOpenGLCanvas |
| | OpenGL Canvas for 3D-visualization of map data. More...
|
| |
| class | Spectrum3DWidget |
| | Widget for 3D-visualization of map data. More...
|
| |
| class | SpectrumAccessOpenMS |
| | An implementation of the OpenSWATH Spectrum Access interface using OpenMS. More...
|
| |
| class | SpectrumAccessOpenMSCached |
| | An implementation of the Spectrum Access interface using on-disk caching. More...
|
| |
| class | SpectrumAccessOpenMSInMemory |
| | An implementation of the OpenSWATH Spectrum Access interface completely in memory. More...
|
| |
| class | SpectrumAccessQuadMZTransforming |
| | A transforming m/z wrapper around spectrum access using a quadratic equation. More...
|
| |
| class | SpectrumAccessSqMass |
| | An implementation of the Spectrum Access interface using SQL files. More...
|
| |
| class | SpectrumAccessTransforming |
| | An abstract base class implementing a transforming wrapper around spectrum access. More...
|
| |
| class | SpectrumAddition |
| | The SpectrumAddition is used to add up individual spectra. More...
|
| |
| class | SpectrumAlignment |
| | Aligns the peaks of two sorted spectra Method 1: Using a banded (width via 'tolerance' parameter) alignment if absolute tolerances are given. Scoring function is the m/z distance between peaks. Intensity does not play a role! More...
|
| |
| class | SpectrumAlignmentDialog |
| | Lets the user select two spectra and set the parameters for the spectrum alignment. More...
|
| |
| class | SpectrumAlignmentScore |
| | Similarity score via spectra alignment. More...
|
| |
| class | SpectrumAnnotator |
| | Annotates spectra from identifications and theoretical spectra or identifications from spectra and theoretical spectra matching with various options. More...
|
| |
| class | SpectrumCanvas |
| | Base class for visualization canvas classes. More...
|
| |
| class | SpectrumCheapDPCorr |
| | SpectrumCheapDPCorr calculates an optimal alignment on stick spectra. More...
|
| |
| class | SpectrumIdentification |
| | Represents a object which can store the information of an analysisXML instance. More...
|
| |
| class | SpectrumLookup |
| | Helper class for looking up spectra based on different attributes. More...
|
| |
| class | SpectrumMetaDataLookup |
| | Helper class for looking up spectrum meta data. More...
|
| |
| class | SpectrumPrecursorComparator |
| | SpectrumPrecursorComparator compares just the parent mass of two spectra. More...
|
| |
| class | SpectrumSettings |
| | Representation of 1D spectrum settings. More...
|
| |
| class | SpectrumSettingsVisualizer |
| | Class that displays all meta information for SpectrumSettings objects. More...
|
| |
| class | SpectrumWidget |
| | Base class for spectrum widgets. More...
|
| |
| class | SplineInterpolatedPeaks |
| | Data structure for spline interpolation of MS1 spectra and chromatograms. More...
|
| |
| class | SplinePackage |
| | fundamental data structure for SplineInterpolatedPeaks More...
|
| |
| class | SqliteConnector |
| | File adapter for Sqlite files. More...
|
| |
| class | SqMassFile |
| | An class that uses on-disk SQLite database to read and write spectra and chromatograms. More...
|
| |
| class | SqrtMower |
| | Scales the intensity of peaks to the sqrt. More...
|
| |
| class | StablePairFinder |
| | This class implements a pair finding algorithm for consensus features. More...
|
| |
| class | SteinScottImproveScore |
| | Similarity score based of Stein & Scott. More...
|
| |
| class | StopWatch |
| | StopWatch Class. More...
|
| |
| class | StreamHandler |
| | Provides a central class to register globally used output streams. Currently supported streams are. More...
|
| |
| class | String |
| | A more convenient string class. More...
|
| |
| class | StringListUtils |
| | Utilities operating on lists of Strings. More...
|
| |
| class | StringUtils |
| |
| class | StringView |
| | StringView provides a non-owning view on an existing string. More...
|
| |
| struct | Summary |
| | Summary of fitting results. More...
|
| |
| class | SuperHirnParameters |
| | SuperHirn parameters singleton class containing all static configuration variables. More...
|
| |
| class | SuperHirnUtil |
| |
| struct | SVMData |
| | Data structure used in SVMWrapper. More...
|
| |
| class | SvmTheoreticalSpectrumGenerator |
| | Simulates MS2 spectra with support vector machines. More...
|
| |
| class | SvmTheoreticalSpectrumGeneratorSet |
| | Loads SvmTheoreticalSpectrumGenerator instances for different charges. More...
|
| |
| class | SvmTheoreticalSpectrumGeneratorTrainer |
| | Train SVM models that are used by SvmTheoreticalSpectrumGenerator. More...
|
| |
| class | SVMWrapper |
| | Serves as a wrapper for the libsvm. More...
|
| |
| class | SVOutStream |
| | Stream class for writing to comma/tab/...-separated values files. More...
|
| |
| class | SwathFile |
| | File adapter for Swath files. More...
|
| |
| class | SwathMapMassCorrection |
| | A class containing correction functions for Swath MS maps. More...
|
| |
| class | SwathWindowLoader |
| | Class to read a file describing the Swath Windows. More...
|
| |
| class | SysInfo |
| | Some functions to get system information. More...
|
| |
| class | Tagger |
| | Calculate sequence tags from m/z values. More...
|
| |
| class | Tagging |
| | Meta information about tagging of a sample e.g. ICAT labeling. More...
|
| |
| class | TaggingVisualizer |
| | Class that displays all meta information of tagging objects. More...
|
| |
| class | TargetedExperiment |
| | A description of a targeted experiment containing precursor and production ions. More...
|
| |
| class | TargetedSpectraExtractor |
| | This class filters, annotates, picks, and scores spectra (e.g., taken from a DDA experiment) based on a target list. More...
|
| |
| class | TextFile |
| | This class provides some basic file handling methods for text files. More...
|
| |
| class | TheoreticalSpectrumGenerationDialog |
| | Dialog which allows to enter an AA sequence and generates a theoretical spectrum for it. More...
|
| |
| class | TheoreticalSpectrumGenerator |
| | Generates theoretical spectra for peptides with various options. More...
|
| |
| class | TheoreticalSpectrumGeneratorXLMS |
| | Generates theoretical spectra for cross-linked peptides. More...
|
| |
| class | ThresholdMower |
| | ThresholdMower removes all peaks below a threshold. More...
|
| |
| class | TIC |
| |
| class | TICFilter |
| | TICFilter calculates TIC. More...
|
| |
| class | TMTElevenPlexQuantitationMethod |
| | TMT 11plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | TMTSixPlexQuantitationMethod |
| | TMT 6plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | TMTSixteenPlexQuantitationMethod |
| | TMT 16plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | TMTTenPlexQuantitationMethod |
| | TMT 10plex quantitation to be used with the IsobaricQuantitation. More...
|
| |
| class | TOFCalibration |
| | This class implements an external calibration for TOF data using external calibrant spectra. More...
|
| |
| class | ToolDescriptionFile |
| | File adapter for ToolDescriptor files. More...
|
| |
| class | ToolHandler |
| |
| class | ToolsDialog |
| | TOPP tool selection dialog. More...
|
| |
| class | TOPPASBase |
| | Main window of the TOPPAS tool. More...
|
| |
| class | TOPPASEdge |
| | An edge representing a data flow in TOPPAS. More...
|
| |
| class | TOPPASInputFileDialog |
| | Dialog which allows to specify an input file. More...
|
| |
| class | TOPPASInputFileListVertex |
| | A vertex representing an input file list. More...
|
| |
| class | TOPPASInputFilesDialog |
| | Dialog which allows to specify a list of input files. More...
|
| |
| class | TOPPASIOMappingDialog |
| | Dialog which allows to configure the input/output parameter mapping of an edge. More...
|
| |
| class | TOPPASLogWindow |
| | QTextEdit implementation with a "clear" button in the context menu. More...
|
| |
| class | TOPPASMergerVertex |
| | A special vertex that allows to merge several inputs. More...
|
| |
| class | TOPPASOutputFileListVertex |
| | A vertex representing an output file list. More...
|
| |
| class | TOPPASOutputFilesDialog |
| | Dialog which allows to specify the directory for the output files. More...
|
| |
| class | TOPPASResource |
| | Represents a data resource for TOPPAS workflows. More...
|
| |
| class | TOPPASResources |
| | A dictionary mapping string keys to lists of TOPPASResource objects. More...
|
| |
| class | TOPPASScene |
| | A container for all visual items of a TOPPAS workflow. More...
|
| |
| class | TOPPASSplitterVertex |
| | A special vertex that allows to split a list of inputs. More...
|
| |
| class | TOPPASTabBar |
| | Convenience tab bar implementation. More...
|
| |
| class | TOPPASToolConfigDialog |
| | TOPP tool configuration dialog. More...
|
| |
| class | TOPPASToolVertex |
| | A vertex representing a TOPP tool. More...
|
| |
| class | TOPPASTreeView |
| | Tree view implementation for the list of TOPP tools. More...
|
| |
| class | TOPPASVertex |
| | The base class of the different vertex classes. More...
|
| |
| class | TOPPASVertexNameDialog |
| | Dialog which allows to change the name of an input/output vertex. More...
|
| |
| class | TOPPASWidget |
| | Widget visualizing and allowing to edit TOPP pipelines. More...
|
| |
| class | TOPPBase |
| | Base class for TOPP applications. More...
|
| |
| class | TOPPOpenSwathBase |
| |
| class | TOPPViewBase |
| | Main window of TOPPView tool. More...
|
| |
| class | TOPPViewIdentificationViewBehavior |
| | Behavior of TOPPView in identification mode. More...
|
| |
| class | TOPPViewOpenDialog |
| | Dataset opening options for TOPPView. More...
|
| |
| class | TOPPViewSpectraViewBehavior |
| | Behavior of TOPPView in spectra view mode. More...
|
| |
| class | TraceFitter |
| | Abstract fitter for RT profile fitting. More...
|
| |
| class | TraMLFile |
| | File adapter for HUPO PSI TraML files. More...
|
| |
| class | TransformationDescription |
| | Generic description of a coordinate transformation. More...
|
| |
| class | TransformationModel |
| | Base class for transformation models. More...
|
| |
| class | TransformationModelBSpline |
| | B-spline (non-linear) model for transformations. More...
|
| |
| class | TransformationModelInterpolated |
| | Interpolation model for transformations. More...
|
| |
| class | TransformationModelLinear |
| | Linear model for transformations. More...
|
| |
| class | TransformationModelLowess |
| | Lowess (non-linear) model for transformations. More...
|
| |
| class | TransformationXMLFile |
| | Used to load and store TransformationXML files. More...
|
| |
| class | TransitionGroupOpenMS |
| | An implementation of the OpenSWATH Transition Group Access interface using OpenMS. More...
|
| |
| class | TransitionPQPFile |
| | This class supports reading and writing of PQP files. More...
|
| |
| class | TransitionTSVFile |
| | This class supports reading and writing of OpenSWATH transition lists. More...
|
| |
| class | TwoDOptimization |
| | This class provides the two-dimensional optimization of the picked peak parameters. More...
|
| |
| class | UnimodXMLFile |
| | Used to load XML files from unimod.org files. More...
|
| |
| class | UniqueIdGenerator |
| | A generator for unique ids. More...
|
| |
| class | UniqueIdIndexer |
| | A base class for random access containers for classes derived from UniqueIdInterface that adds functionality to convert a unique id into an index into the container. More...
|
| |
| class | UniqueIdInterface |
| | A base class defining a common interface for all classes having a unique id. More...
|
| |
| class | UnnormalizedComparator |
| | Exception thrown if clustering is attempted without a normalized compare functor. More...
|
| |
| class | UpdateCheck |
| | Helper Functions to perform an update query to the OpenMS REST server. More...
|
| |
| struct | ValueTrait |
| | Trait for MatchedIterator to find pairs with a certain distance, which is computed directly on the value_type of the container. More...
|
| |
| struct | VecLowPrecision |
| |
| class | VersionInfo |
| | Version information class. More...
|
| |
| class | WeightWrapper |
| | Encapsulated weight queries to simplify mono vs average weight computation. More...
|
| |
| class | WindowMower |
| | WindowMower augments the highest peaks in a sliding or jumping window. More...
|
| |
| class | XFDRAlgorithm |
| |
| class | XMassFile |
| | File adapter for 'XMass Analysis (fid)' files. More...
|
| |
| class | XMLValidator |
| | Validator for XML files. More...
|
| |
| class | XQuestResultXMLFile |
| | Used to load and store xQuest result files. More...
|
| |
| class | XQuestScores |
| | An implementation of the scores for cross-link identification from the xQuest algorithm (O. Rinner et al., 2008, "Identification of cross-linked peptides from large sequence databases") More...
|
| |
| class | XTandemInfile |
| | XTandem input file. More...
|
| |
| class | XTandemXMLFile |
| | Used to load XTandemXML files. More...
|
| |
| class | ZhangSimilarityScore |
| | Similarity score of Zhang. More...
|
| |
| class | ZlibCompression |
| | Compresses and uncompresses data using zlib. More...
|
| |
|
| std::ostream & | operator<< (std::ostream &os, const AccurateMassSearchResult &amsr) |
| |
| template<> |
| String | ChromatogramExtractor::extract_id_< OpenSwath::LightTargetedExperiment > (OpenSwath::LightTargetedExperiment &transition_exp_used, const String &id, int &prec_charge) |
| |
| template<> |
| String | ChromatogramExtractor::extract_id_< OpenMS::TargetedExperiment > (OpenMS::TargetedExperiment &transition_exp_used, const String &id, int &prec_charge) |
| |
| std::ostream & | operator<< (std::ostream &os, const PSLPFormulation::IndexTriple &triple) |
| |
| std::ostream & | operator<< (std::ostream &os, const AASequence &peptide) |
| |
| std::istream & | operator>> (std::istream &os, const AASequence &peptide) |
| |
| std::ostream & | operator<< (std::ostream &os, const DigestionEnzyme &enzyme) |
| |
| std::ostream & | operator<< (std::ostream &os, const DigestionEnzymeProtein &enzyme) |
| |
| std::ostream & | operator<< (std::ostream &, const Element &) |
| |
| std::ostream & | operator<< (std::ostream &os, const EmpiricalFormula &formula) |
| |
| std::ostream & | operator<< (std::ostream &os, const Residue &residue) |
| |
| std::ostream & | operator<< (std::ostream &os, const Ribonucleotide &ribo) |
| | Stream output operator. More...
|
| |
| bool | compareBinaryTreeNode (const BinaryTreeNode &x, const BinaryTreeNode &y) |
| | returns the value of (x.distance < y.distance) for use with sort More...
|
| |
| template<UInt N, typename T > |
| std::size_t | hash_value (const DPosition< N, T > &b) |
| |
| std::ostream & | operator<< (std::ostream &os, const Exception::BaseException &e) |
| | Output operator for exceptions. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, LogConfigHandler const &lch) |
| | Overload for the insertion operator (operator<<) to have a formatted output of the LogConfigHandler. More...
|
| |
| template<typename FloatingPointType > |
| const PrecisionWrapper< FloatingPointType > | precisionWrapper (const FloatingPointType rhs) |
| | Wrapper function that sets the appropriate precision for output temporarily. The original precision is restored afterwards so that no side effects remain. This is a "make"-function that deduces the typename FloatingPointType from its argument and returns a PrecisionWrapper<FloatingPointType>. More...
|
| |
| template<typename FloatingPointType > |
| std::ostream & | operator<< (std::ostream &os, const PrecisionWrapper< FloatingPointType > &rhs) |
| | Output operator for a PrecisionWrapper. Specializations are defined for float, double, long double. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, StreamHandler const &stream_handler) |
| | Overload for the insertion operator (operator<<) to have a formatted output of the StreamHandler. More...
|
| |
| template<typename Type > |
| std::string | typeAsString (const Type &=Type()) |
| | Returns the Type as as std::string. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ChargePair &cons) |
| | Print the contents of a ChargePair to a stream. More...
|
| |
| template<UInt D> |
| std::ostream & | operator<< (std::ostream &os, const DBoundingBox< D > &bounding_box) |
| | Print the contents to a stream. More...
|
| |
| template<typename Value > |
| std::ostream & | operator<< (std::ostream &os, const DistanceMatrix< Value > &matrix) |
| | Print the contents to a stream. More...
|
| |
| template<UInt D, typename TCoordinateType > |
| DPosition< D, TCoordinateType > | operator* (DPosition< D, TCoordinateType > position, typename DPosition< D, TCoordinateType >::CoordinateType scalar) |
| | Scalar multiplication (a bit inefficient) More...
|
| |
| template<UInt D, typename TCoordinateType > |
| DPosition< D, TCoordinateType > | operator* (typename DPosition< D, TCoordinateType >::CoordinateType scalar, DPosition< D, TCoordinateType > position) |
| | Scalar multiplication (a bit inefficient) More...
|
| |
| template<UInt D, typename TCoordinateType > |
| DPosition< D, TCoordinateType > | operator/ (DPosition< D, TCoordinateType > position, typename DPosition< D, TCoordinateType >::CoordinateType scalar) |
| | Scalar multiplication (a bit inefficient) More...
|
| |
| template<UInt D, typename TCoordinateType > |
| std::ostream & | operator<< (std::ostream &os, const DPosition< D, TCoordinateType > &pos) |
| | Print the contents to a stream. More...
|
| |
| template<UInt D> |
| std::ostream & | operator<< (std::ostream &os, const DRange< D > &area) |
| | Print the contents to a stream. More...
|
| |
| template<typename T > |
| std::ostream & | operator<< (std::ostream &os, const std::vector< T > &v) |
| | Output stream operator for std::vectors. More...
|
| |
| template<typename T > |
| std::ostream & | operator<< (std::ostream &os, const VecLowPrecision< T > &val) |
| |
| template<typename TString > |
| std::vector< String > & | operator<< (std::vector< String > &sl, const TString &string) |
| | Operator for appending entries with less code. More...
|
| |
| template<typename Value > |
| std::ostream & | operator<< (std::ostream &os, const Matrix< Value > &matrix) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const Param ¶m) |
| | Output of Param to a stream. More...
|
| |
| ::size_t | hash_value (OpenMS::String const &s) |
| |
| static EigenMatrixXdPtr | convertOpenMSMatrix2EigenMatrixXd (const Matrix< double > &m) |
| |
| static bool | matrixIsIdentityMatrix (const Matrix< double > &channel_frequency) |
| |
| UInt32 | endianize32 (const UInt32 &n) |
| | Endianizes a 32 bit type from big endian to little endian and vice versa. More...
|
| |
| UInt64 | endianize64 (const UInt64 &n) |
| | Endianizes a 64 bit type from big endian to little endian and vice versa. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ControlledVocabulary &cv) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ChromatogramPeak &point) |
| | Print the contents to a stream. More...
|
| |
| template<class Cmp > |
| PointerComparator< Cmp > | pointerComparator (Cmp const &cmp) |
| | Make-function to create a PointerComparator from another comparator without the need to specify the template arguments. More...
|
| |
| template<class Cmp > |
| ReverseComparator< Cmp > | reverseComparator (Cmp const &cmp) |
| | Make-function to create a ReverseComparator from another comparator without the need to specify the template arguments. More...
|
| |
| template<typename Cmp1 , typename Cmp2 > |
| LexicographicComparator< Cmp1, Cmp2 > | lexicographicComparator (Cmp1 const &cmp1, Cmp2 const &cmp2) |
| | Make-function to create a LexicographicComparator from two other comparators without the need to specify the template arguments. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ConsensusFeature &cons) |
| | Print the contents of a ConsensusFeature to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ConsensusMap &cons_map) |
| | Print the contents of a ConsensusMap to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const FeatureHandle &cons) |
| | Print the contents of a FeatureHandle to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const AnnotationStatistics &ann) |
| | Print content of an AnnotationStatistics object to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const FeatureMap &map) |
| |
| std::ostream & | operator<< (std::ostream &os, const MSChromatogram &chrom) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const MSExperiment &exp) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const MSSpectrum &spec) |
| |
| std::ostream & | operator<< (std::ostream &os, const Peak1D &point) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const Peak2D &point) |
| | Print the contents to a stream. More...
|
| |
| template<class DataArrayT > |
| DataArrayT::iterator | getDataArrayByName (DataArrayT &a, const String &name) |
| | Helper functions for MSSpectrum and MSChromatogram. More...
|
| |
| template<class DataArrayT > |
| DataArrayT::const_iterator | getDataArrayByName (const DataArrayT &a, const String &name) |
| |
| template<typename PeakContainerT > |
| void | removePeaks (PeakContainerT &p, const double pos_start, const double pos_end, const bool ignore_data_arrays=false) |
| | remove all peaks EXCEPT in the given range More...
|
| |
| template<typename PeakContainerT > |
| void | subtractMinimumIntensity (PeakContainerT &p) |
| |
| template<typename PeakContainerT > |
| void | makePeakPositionUnique (PeakContainerT &p, const IntensityAveragingMethod m=IntensityAveragingMethod::MEDIAN) |
| | Make peak positions unique. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ChromatogramSettings &spec) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const ExperimentalSettings &exp) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const SpectrumSettings &spec) |
| | Print the contents to a stream. More...
|
| |
| std::ostream & | operator<< (std::ostream &os, const QCBase::Status &stat) |
| |
| template<typename PeakType > |
| bool | intensityComparator (const PeakType &a, const PeakType &b) |
| |
| template<typename PeakType > |
| bool | intensityAscendingComparator (const PeakType &a, const PeakType &b) |
| |
| template<typename PeakType > |
| bool | intensityPointerComparator (PeakType *a, PeakType *b) |
| |
| template<typename PeakType > |
| bool | positionComparator (const PeakType &a, const PeakType &b) |
| |
| bool | operator< (const MultiplexDeltaMasses &dm1, const MultiplexDeltaMasses &dm2) |
| |
| bool | sort_peaks_by_intensity (const PeakCandidate &a, const PeakCandidate &b) |
| |
| std::ostream & | operator<< (std::ostream &, CentroidData &) |
| |
| std::ostream & | operator<< (std::ostream &, CentroidPeak &) |
| |
| std::ostream & | operator<< (std::ostream &, DeconvPeak &) |
| |
| SUPERHIRN_DLLAPI std::ostream & | operator<< (std::ostream &, Deisotoper &) |
| |
| std::ostream & | operator<< (std::ostream &out, RawData &data) |
| |
| std::ostream & | operator<< (std::ostream &os, const LayerData &rhs) |
| | Print the contents to a stream. More...
|
| |
| IMType | determineIMType (const MSExperiment &exp) |
| |
| void | processDriftTimeStack (const std::vector< MSSpectrum > &stack, std::vector< MSSpectrum > &result) |
| | Process a stack of drift time spectra. More...
|
| |
| void | expandIMSpectrum (const MSSpectrum &tmps, std::vector< MSSpectrum > &result) |
| | Expands a single MSSpectrum (single frame) into individual ion mobility spectrum. More...
|
| |
| void | collapseIMSpectrum (const MSExperiment &exp, std::vector< MSSpectrum > &result) |
| | Collapses multiple IM spectra from the same frame into a single MSSpectrum. More...
|
| |
|
These functions are provided to unify the handling of this issue throughout OpenMS. (So please don't use ad-hoc numbers ;-) )
If you want to avoid side effects you can use precisionWrapper() to write a floating point number with appropriate precision; in this case the original state of the stream is automatically restored afterwards. See precisionWrapper() for details.
In practice, the number of decimal digits that the type can represent without loss of precision are 6 digits for single precision and 15 digits for double precision. We have 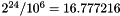 and and 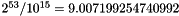 , so rounding will remove the remaining difference. , so rounding will remove the remaining difference.
Example: #define NUMBER 12345.67890123456789012345678901
std::cout << NUMBER << '\n';
double d = NUMBER;
std::cout << writtenDigits<double>(0.0) << ": " << d << '\n';
float r = NUMBER;
long double l = NUMBER;
double x = 88.99;
std::cout.precision(15);
std::cout << "15: " << x << '\n';
std::cout.precision(16);
std::cout << "16: " << x << '\n';
|
| template<typename FloatingPointType > |
| constexpr Int | writtenDigits (const FloatingPointType &=FloatingPointType()) |
| | Number of digits commonly used for writing a floating point type (a.k.a. precision). Specializations are defined for float, double, long double. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< float > (const float &) |
| | Number of digits commonly used for writing a float (a.k.a. precision). More...
|
| |
| template<> |
| constexpr Int | writtenDigits< double > (const double &) |
| | Number of digits commonly used for writing a double (a.k.a. precision). More...
|
| |
| template<> |
| constexpr Int | writtenDigits< int > (const int &) |
| | We do not want to bother people who unintentionally provide an int argument to this. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< unsigned int > (const unsigned int &) |
| | We do not want to bother people who unintentionally provide an unsigned int argument to this. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< long int > (const long int &) |
| | We do not want to bother people who unintentionally provide a long int argument to this. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< unsigned long int > (const unsigned long int &) |
| | We do not want to bother people who unintentionally provide an unsigned long int argument to this. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< DataValue > (const DataValue &) |
| | DataValue will be printed like double. More...
|
| |
| template<> |
| constexpr Int | writtenDigits< long double > (const long double &) |
| | Number of digits commonly used for writing a long double (a.k.a. precision). ... More...
|
| |


 1.8.16
1.8.16